Machine Learning Model Deployment: How to Deploy a Machine Learning Model to Production (Step-by-Step)
Machine learning model deployment is where most ML projects fail.
The model works in a notebook. The demo looks great. But taking it into production is a completely different challenge.
Series: Production ML Engineering — Article 02 of 15 (Cluster 1: Foundation & Pipeline)
Before you read this: This article is part of a 15-part series on building production-grade ML systems. If you have not read the series hub yet, start with the Production ML Engineering guide — it maps out the five pillars every production system rests on and explains where this article fits in the bigger picture.
Every time I talk to an ML engineer who has been in the field long enough, the story is remarkably consistent: the model worked beautifully in the notebook. Stakeholders were impressed by the demo. Then someone asked “how do we put this in front of real users?” — and the next three months became an unplanned education in distributed systems, container orchestration, and things nobody warns you about in any machine learning course.
I have been on both sides of that experience. The first time I tried to move a trained sklearn model into a service that real traffic could hit, I did what most people do: wrapped it in a Flask endpoint, ran it on a VM, and called it shipped. It ran fine for about six days. Then a dependency updated itself on the server, a colleague restarted the process without the right environment variables loaded, and the service quietly died at 2am on a Sunday. No alerts. No rollback. Just a dead endpoint and a very unpleasant Monday morning.
What I did not understand yet was that deployment is not an event. It is a system. Getting a model to respond to one request in a local terminal is a solved problem — any tutorial will get you there. Getting it to handle variable load, recover gracefully from failures, integrate into a CI/CD pipeline, and deploy new versions without downtime is a completely different engineering discipline.
This article documents what that system actually looks like, built from scratch, with every design decision explained. By the end, you will have a working ML service with a FastAPI REST API, a two-stage Docker build, a full test suite, and a GitHub Actions pipeline that tests, builds, and deploys automatically. Everything is code-first. Nothing is hand-wavy.
Complete Code: https://github.com/Emmimal/ml-service/
Why Notebook-to-Production Breaks Most Teams
The gap is not about model quality. I have seen excellent models fail to ship and mediocre ones run in production for years. The gap is about what the notebook hides from you.
A notebook assumes a static environment. You have one version of scikit-learn, one version of numpy, one fixed dataset, and you run top-to-bottom whenever you want results. The state lives in your kernel. Restart the kernel and you rebuild it by running the cells again. That is fine for research. It is a reliability disaster for a service that needs to recover automatically after a crash, serve multiple concurrent requests, and produce the exact same transformation logic at inference time that it used at training time.
Here is what silently breaks when you move from notebook to production:
Training-serving skew. You fit a StandardScaler in the notebook and save the model weights. But at inference time, you compute the scaling parameters again from scratch — or you forget to apply scaling at all. The model sees inputs on a completely different distribution than it was trained on. It still runs. It still returns predictions. They are just wrong, and nothing will tell you.
Environment drift. Your notebook ran on Python 3.10 with scikit-learn 1.3.2. The server runs Python 3.8 with scikit-learn 1.0 because that was what the sysadmin had available. Behaviour diverges in subtle ways that surface as incorrect predictions rather than obvious errors.
No recovery path. When the process crashes — and it will — there is no restart policy, no health check, no mechanism to detect the outage and route traffic away from the broken instance.
Manual deployment. The steps to get a new model version into production live in someone’s head, or in a Slack message from three months ago. The first time that person is unavailable during a deployment, the team is stuck.
The five-step workflow that follows addresses all of these problems in sequence. Each step is designed to eliminate a specific failure mode.
The Project We Are Building
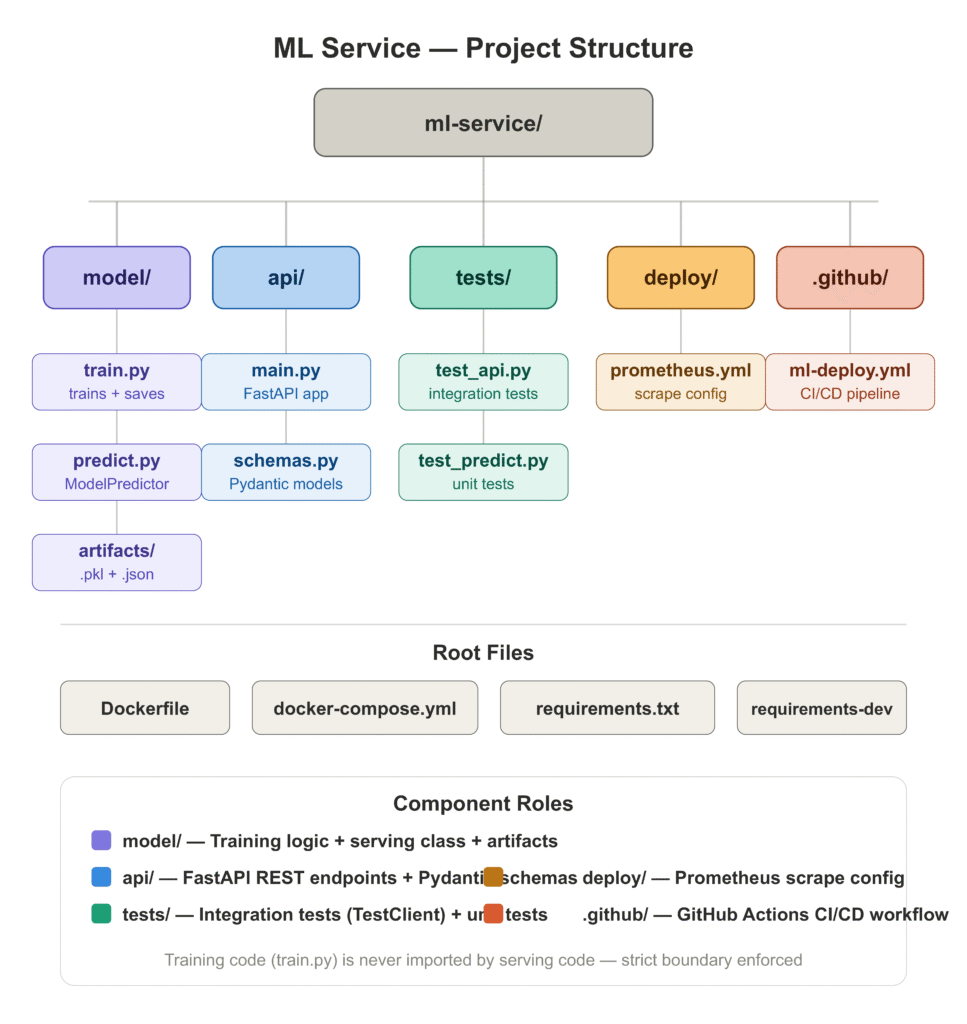
Before walking through each step, here is what the completed project looks like:
ml-service/
├── model/
│ ├── train.py # trains and persists artifacts
│ ├── predict.py # ModelPredictor class
│ └── artifacts/ # model.pkl, preprocessor.pkl, metadata.json
├── api/
│ ├── main.py # FastAPI application
│ └── schemas.py # Pydantic request/response models
├── tests/
│ ├── test_api.py # integration tests (TestClient)
│ └── test_predict.py # unit tests (ModelPredictor)
├── deploy/
│ └── prometheus.yml # Prometheus scrape config
├── .github/
│ └── workflows/
│ └── ml-deploy.yml # CI/CD pipeline
├── Dockerfile
├── docker-compose.yml
├── requirements.txt
└── requirements-dev.txtWe are using a GradientBoostingClassifier trained on the UCI Breast Cancer dataset [1] — a clean, well-understood binary classification problem that lets us focus on the deployment system rather than on feature engineering. The same architecture applies directly to any sklearn-compatible model.
Step 1: Structure Your ML Code for Deployment
The most important decision you make before writing a single line of serving code is how you separate your concerns. Notebooks do not enforce this — everything lives in one file and state bleeds everywhere. Production services need explicit boundaries.
The two boundaries that matter most are:
Training from serving. The code that fits the model should never be imported by the code that serves predictions. If it is, you risk accidentally retrigger training at inference time, you make the serving container larger than it needs to be, and you create a dependency between two systems that should be able to evolve independently.
The preprocessor from the model. This is the one that surprises people. When you use an sklearn Pipeline that bundles your scaler and your model together, it feels clean — one .fit(), one .predict(). But it makes debugging harder in production. If predictions are wrong, you want to be able to inspect what the scaler did independently of what the model did. Saving them as separate artifacts gives you that.
Training: model/train.py
"""
model/train.py
--------------
Train a GradientBoostingClassifier on the UCI Breast Cancer dataset and
persist the preprocessor and model as separate joblib artifacts.
Keeping them separate matters in production: you can inspect, benchmark,
or swap the preprocessor without touching the model weights, and you get
an exact record of what transformation the serving code must reproduce.
Usage:
python -m model.train
Output:
model/artifacts/preprocessor.pkl
model/artifacts/model.pkl
model/artifacts/metadata.json
"""
import json
import logging
import time
from pathlib import Path
import joblib
import numpy as np
from sklearn.datasets import load_breast_cancer
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.metrics import (
classification_report,
roc_auc_score,
average_precision_score,
)
from sklearn.model_selection import StratifiedKFold, cross_val_score, train_test_split
from sklearn.preprocessing import StandardScaler
logging.basicConfig(
level=logging.INFO,
format="%(asctime)s %(levelname)-8s %(message)s",
datefmt="%Y-%m-%d %H:%M:%S",
)
logger = logging.getLogger(__name__)
ARTIFACT_DIR = Path(__file__).parent / "artifacts"
ARTIFACT_DIR.mkdir(parents=True, exist_ok=True)
MODEL_CONFIG = {
"n_estimators": 200,
"learning_rate": 0.05,
"max_depth": 4,
"subsample": 0.8,
"random_state": 42,
}
RANDOM_STATE = 42
TEST_SIZE = 0.20
def evaluate(model, X: np.ndarray, y: np.ndarray, split_name: str) -> dict:
"""Return a dict of evaluation metrics for a given split."""
y_pred = model.predict(X)
y_proba = model.predict_proba(X)[:, 1]
report = classification_report(y, y_pred, output_dict=True)
metrics = {
"split": split_name,
"accuracy": round(report["accuracy"], 4),
"precision_macro": round(report["macro avg"]["precision"], 4),
"recall_macro": round(report["macro avg"]["recall"], 4),
"f1_macro": round(report["macro avg"]["f1-score"], 4),
"roc_auc": round(roc_auc_score(y, y_proba), 4),
"avg_precision": round(average_precision_score(y, y_proba), 4),
}
logger.info("[%s] %s", split_name, metrics)
return metrics
def train_and_save() -> None:
logger.info("Loading breast cancer dataset...")
data = load_breast_cancer()
X, y = data.data, data.target
feature_names = list(data.feature_names)
logger.info("Dataset shape: %s | Class balance: %s", X.shape, np.bincount(y))
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=TEST_SIZE, random_state=RANDOM_STATE, stratify=y
)
# StandardScaler is fit on training data only.
# This is the exact object that must be used at serve time.
preprocessor = StandardScaler()
X_train_scaled = preprocessor.fit_transform(X_train)
X_test_scaled = preprocessor.transform(X_test)
logger.info("Running 5-fold stratified cross-validation...")
cv = StratifiedKFold(n_splits=5, shuffle=True, random_state=RANDOM_STATE)
cv_scores = cross_val_score(
GradientBoostingClassifier(**MODEL_CONFIG),
X_train_scaled,
y_train,
cv=cv,
scoring="roc_auc",
n_jobs=-1,
)
logger.info("CV ROC-AUC: %.4f ± %.4f", cv_scores.mean(), cv_scores.std())
logger.info("Training final model...")
t0 = time.perf_counter()
model = GradientBoostingClassifier(**MODEL_CONFIG)
model.fit(X_train_scaled, y_train)
train_duration_s = round(time.perf_counter() - t0, 2)
logger.info("Training completed in %.2fs", train_duration_s)
train_metrics = evaluate(model, X_train_scaled, y_train, "train")
test_metrics = evaluate(model, X_test_scaled, y_test, "test")
print("\n── Held-out Test Set ──────────────────────────────")
print(classification_report(y_test, model.predict(X_test_scaled),
target_names=data.target_names))
joblib.dump(preprocessor, ARTIFACT_DIR / "preprocessor.pkl")
joblib.dump(model, ARTIFACT_DIR / "model.pkl")
logger.info("Artifacts saved to %s", ARTIFACT_DIR)
metadata = {
"model_class": type(model).__name__,
"model_config": MODEL_CONFIG,
"feature_names": feature_names,
"n_features": len(feature_names),
"dataset": "sklearn.datasets.load_breast_cancer",
"train_samples": len(X_train),
"test_samples": len(X_test),
"train_duration_s": train_duration_s,
"cv_roc_auc_mean": round(float(cv_scores.mean()), 4),
"cv_roc_auc_std": round(float(cv_scores.std()), 4),
"train_metrics": train_metrics,
"test_metrics": test_metrics,
}
with open(ARTIFACT_DIR / "metadata.json", "w") as f:
json.dump(metadata, f, indent=2)
logger.info("Metadata written to %s", ARTIFACT_DIR / "metadata.json")
if __name__ == "__main__":
train_and_save()Running python -m model.train produced this output in my environment:
2026-04-21 00:48:31 INFO Loading breast cancer dataset...
2026-04-21 00:48:31 INFO Dataset shape: (569, 30) | Class balance: [212 357]
2026-04-21 00:48:31 INFO Running 5-fold stratified cross-validation...
2026-04-21 00:48:43 INFO CV ROC-AUC: 0.9912 ± 0.0063
2026-04-21 00:48:43 INFO Training final model...
2026-04-21 00:48:44 INFO Training completed in 1.29s
2026-04-21 00:48:44 INFO [train] {'accuracy': 1.0, 'roc_auc': 1.0, ...}
2026-04-21 00:48:44 INFO [test] {'accuracy': 0.9561, 'roc_auc': 0.9917, ...}
── Held-out Test Set ──────────────────────────────
precision recall f1-score support
malignant 0.97 0.90 0.94 42
benign 0.95 0.99 0.97 72
accuracy 0.96 114
macro avg 0.96 0.95 0.95 114
weighted avg 0.96 0.96 0.96 114Strong CV ROC-AUC (0.9912), no sign of data leakage, clean test set performance. The model is worth deploying.
Serving: model/predict.py
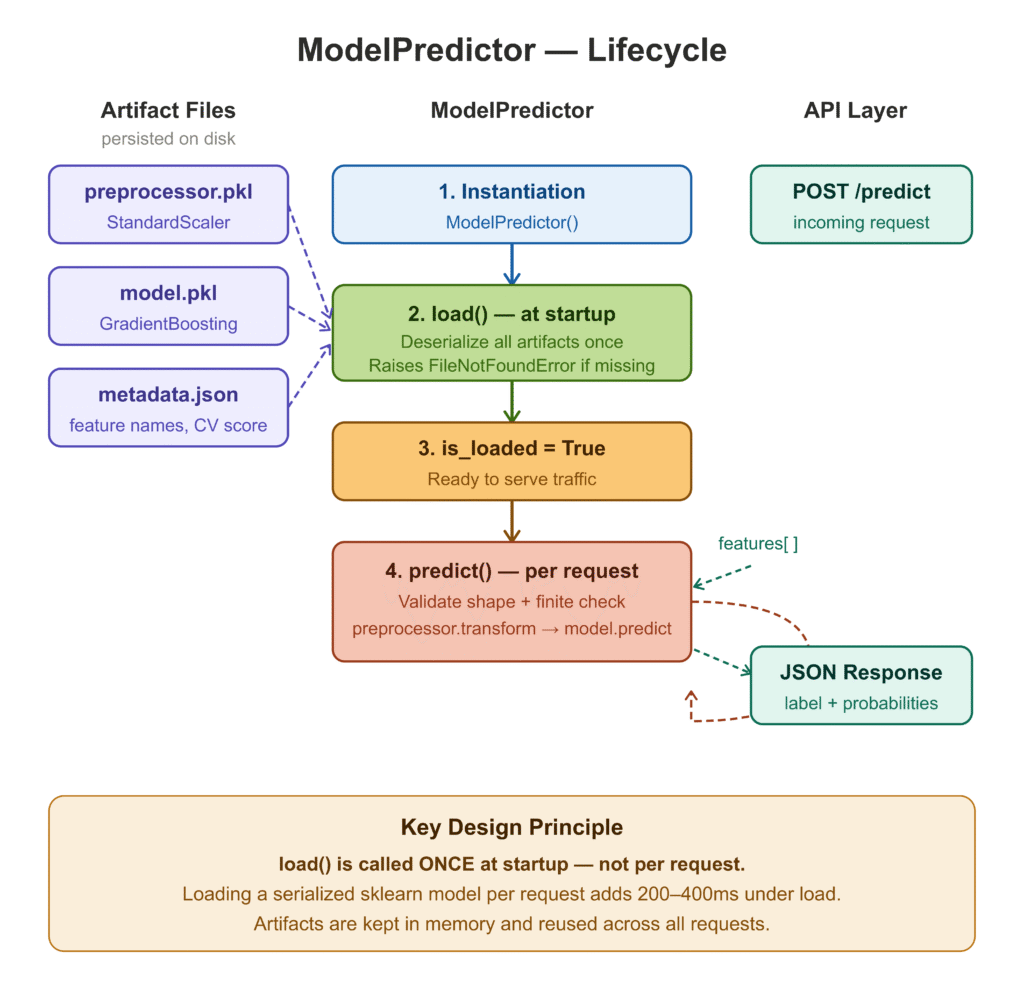
The ModelPredictor class does one thing: load artifacts once at startup and serve predictions on every subsequent call. The distinction between these two operations is the single most important performance decision in ML serving — loading a serialized sklearn model on every request can add 200–400ms under load, even for lightweight models.
"""
model/predict.py
----------------
Wraps artifact loading and inference in a single, reusable class.
Design decisions:
1. Artifacts are loaded once at instantiation, not per request.
2. The preprocessor is kept separate from the model — same as train.py.
This makes training-serving skew immediately visible as an artifact diff.
3. Probabilities are always returned alongside the label so downstream
systems can apply their own decision thresholds.
"""
import json
import logging
from pathlib import Path
from typing import List
import joblib
import numpy as np
logger = logging.getLogger(__name__)
ARTIFACT_DIR = Path(__file__).parent / "artifacts"
class ModelPredictor:
def __init__(self, artifact_dir: Path = ARTIFACT_DIR) -> None:
self._artifact_dir = artifact_dir
self._preprocessor = None
self._model = None
self._metadata: dict = {}
def load(self) -> None:
"""
Deserialize preprocessor, model, and metadata from disk.
Raises FileNotFoundError at startup rather than silently at
the first prediction request.
"""
preprocessor_path = self._artifact_dir / "preprocessor.pkl"
model_path = self._artifact_dir / "model.pkl"
metadata_path = self._artifact_dir / "metadata.json"
for path in (preprocessor_path, model_path):
if not path.exists():
raise FileNotFoundError(
f"Artifact not found: {path}\n"
"Run `python -m model.train` to generate artifacts."
)
self._preprocessor = joblib.load(preprocessor_path)
self._model = joblib.load(model_path)
if metadata_path.exists():
with open(metadata_path) as f:
self._metadata = json.load(f)
logger.info(
"Loaded %s | expects %d features | CV ROC-AUC %.4f",
self._metadata.get("model_class", "model"),
self._metadata.get("n_features", "?"),
self._metadata.get("cv_roc_auc_mean", 0.0),
)
@property
def is_loaded(self) -> bool:
return self._model is not None and self._preprocessor is not None
@property
def expected_n_features(self) -> int:
return self._metadata.get("n_features", 30)
@property
def feature_names(self) -> List[str]:
return self._metadata.get("feature_names", [])
def predict(self, features: List[float]) -> dict:
if not self.is_loaded:
raise RuntimeError("Call .load() before .predict().")
arr = np.array(features, dtype=np.float64).reshape(1, -1)
if arr.shape[1] != self.expected_n_features:
raise ValueError(
f"Expected {self.expected_n_features} features, "
f"got {arr.shape[1]}."
)
if not np.isfinite(arr).all():
raise ValueError("Feature vector contains NaN or infinite values.")
scaled = self._preprocessor.transform(arr)
label = int(self._model.predict(scaled)[0])
proba = self._model.predict_proba(scaled)[0]
return {
"label": label,
"probabilities": {
"class_0": round(float(proba[0]), 6),
"class_1": round(float(proba[1]), 6),
},
"n_features_in": arr.shape[1],
}
def predict_batch(self, batch: List[List[float]]) -> List[dict]:
"""
Batch predictions. Amortizes preprocessing overhead —
significantly faster than N sequential .predict() calls.
"""
if not self.is_loaded:
raise RuntimeError("Call .load() before .predict_batch().")
arr = np.array(batch, dtype=np.float64)
if arr.ndim != 2 or arr.shape[1] != self.expected_n_features:
raise ValueError(
f"Batch must have shape (N, {self.expected_n_features}), "
f"got {arr.shape}."
)
scaled = self._preprocessor.transform(arr)
labels = self._model.predict(scaled).tolist()
probas = self._model.predict_proba(scaled).tolist()
return [
{
"label": int(labels[i]),
"probabilities": {
"class_0": round(probas[i][0], 6),
"class_1": round(probas[i][1], 6),
},
}
for i in range(len(labels))
]Notice the three validators in predict(): shape check, finite value check, and the loaded-state guard. All three happen before the preprocessor is touched. Boundary validation is always cheaper than debugging a silent failure three levels down the call stack.
Step 2: Containerize Your Model with Docker
Containerization solves the environment drift problem. Once your service runs correctly inside a Docker image, it will run correctly everywhere that image is deployed — your laptop, a CI runner, a cloud VM, a Kubernetes pod. The container is the environment.
There are two patterns I have seen in production. The first is a single-stage build: install everything into the base image, copy your code, done. Simple to write, easy to understand, and produces an image that is often 1–2 GB because it contains pip, wheel, build tools, and other things the running service never needs. The second is a multi-stage build.
Two-Stage Dockerfile
# ── Stage 1: dependency builder ───────────────────────────────────────────────
# Installing dependencies in a separate stage means they are cached as long
# as requirements.txt does not change — even if application code does.
# This cuts rebuild time from ~60s to < 5s on a warm cache.
FROM python:3.11-slim AS builder
WORKDIR /build
COPY requirements.txt .
RUN pip install --no-cache-dir --prefix=/install -r requirements.txt
# ── Stage 2: runtime image ────────────────────────────────────────────────────
# The runtime image contains no compiler, no pip cache, no build tools.
# Final image size: ~300 MB vs ~800 MB for a single-stage build.
FROM python:3.11-slim AS runtime
# Non-root user. Many container registries and security scanners
# fail images that run as root by default.
RUN groupadd --gid 1001 mluser && \
useradd --uid 1001 --gid mluser --shell /bin/bash --create-home mluser
WORKDIR /app
# Copy compiled packages from builder stage.
COPY --from=builder /install /usr/local
# Copy application source.
COPY api/ ./api/
COPY model/ ./model/
ARG MODEL_VERSION=1.0.0
ENV MODEL_VERSION=${MODEL_VERSION} \
PYTHONDONTWRITEBYTECODE=1 \
PYTHONUNBUFFERED=1 \
PYTHONPATH=/app
# Docker will mark the container unhealthy if this probe fails.
HEALTHCHECK --interval=30s --timeout=5s --start-period=15s --retries=3 \
CMD python -c \
"import urllib.request; urllib.request.urlopen('http://localhost:8000/health')"
USER mluser
EXPOSE 8000
# Exec form so SIGTERM reaches uvicorn directly, not a shell process.
CMD ["uvicorn", "api.main:app", \
"--host", "0.0.0.0", \
"--port", "8000", \
"--workers", "2", \
"--log-level", "info"]Four decisions worth calling out explicitly:
Multi-stage build. The builder stage installs dependencies into /install. The runtime stage copies only those compiled packages — no pip, no wheel, no gcc. The final image is roughly 60% smaller than a single-stage build using the same base image.
Non-root user. mluser with UID 1001 runs the process. This is not paranoia — it is standard practice that many Kubernetes admission controllers and container registries enforce.
HEALTHCHECK. Docker uses this to mark containers healthy or unhealthy in docker ps output and in Swarm-based orchestration. Kubernetes ignores HEALTHCHECK and uses its own liveness/readiness probes, but the habit of writing it is worth keeping.
Exec-form CMD. ["uvicorn", ...] instead of "uvicorn ...". In shell form, Docker wraps the command in /bin/sh -c, which means SIGTERM on container shutdown goes to the shell, not to uvicorn. The process gets SIGKILLed after the grace period expires. Exec form routes the signal directly, allowing uvicorn to finish in-flight requests before exiting.
Local Docker Compose
For local development, Docker Compose wires up the API alongside Prometheus and Grafana so you can observe the service without a cloud account:
version: "3.9"
services:
api:
build:
context: .
target: runtime
args:
MODEL_VERSION: "1.0.0-dev"
image: ml-inference-api:dev
container_name: ml_api
environment:
- MODEL_VERSION=1.0.0-dev
- PYTHONUNBUFFERED=1
ports:
- "8000:8000"
volumes:
- ./api:/app/api:ro
- ./model:/app/model:ro
healthcheck:
test: ["CMD", "python", "-c",
"import urllib.request; urllib.request.urlopen('http://localhost:8000/health')"]
interval: 30s
timeout: 5s
retries: 3
start_period: 15s
restart: unless-stopped
networks:
- ml_net
prometheus:
image: prom/prometheus:v2.53.0
container_name: ml_prometheus
ports:
- "9090:9090"
volumes:
- ./deploy/prometheus.yml:/etc/prometheus/prometheus.yml:ro
- prometheus_data:/prometheus
networks:
- ml_net
grafana:
image: grafana/grafana:11.1.0
container_name: ml_grafana
ports:
- "3000:3000"
environment:
- GF_SECURITY_ADMIN_PASSWORD=admin
- GF_USERS_ALLOW_SIGN_UP=false
volumes:
- grafana_data:/var/lib/grafana
depends_on:
- prometheus
networks:
- ml_net
volumes:
prometheus_data:
grafana_data:
networks:
ml_net:
driver: bridge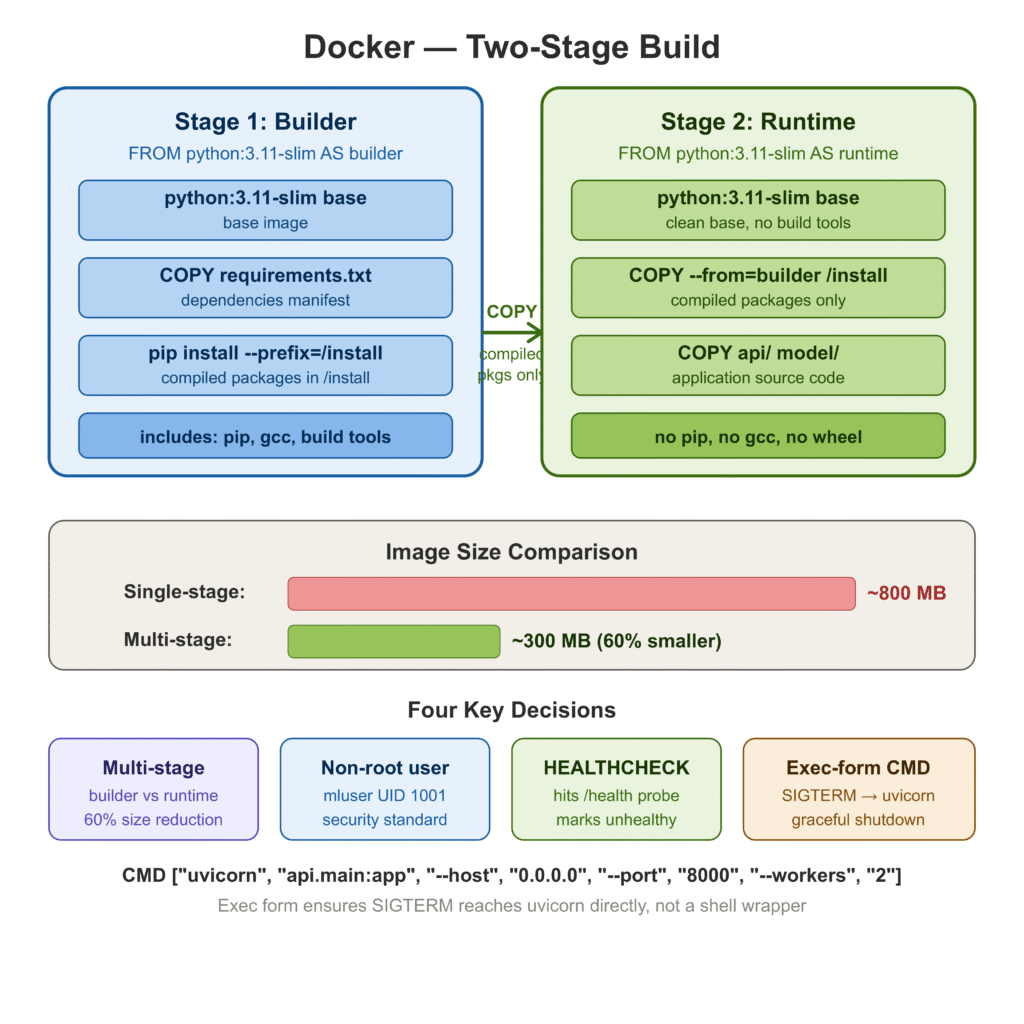
Step 3: Build a REST API with FastAPI
FastAPI [2] is the current standard for ML inference APIs in Python. The reasons practitioners have converged on it are practical: automatic OpenAPI documentation generated from your Python type hints, request validation via Pydantic [3] with no extra work, async support, and performance that comfortably beats Flask on standard benchmarks.
The two files that matter are api/schemas.py (the data contracts) and api/main.py (the application).
Schemas: api/schemas.py
Defining schemas separately from the application logic is a habit that pays off when your API surface grows. Each schema is a Pydantic model. Pydantic handles validation, serialization, and OpenAPI schema generation — you get all three from one class definition.
from typing import Dict, List, Optional
from pydantic import BaseModel, Field, field_validator, model_validator
class PredictRequest(BaseModel):
"""Single-sample inference request."""
features: List[float] = Field(
...,
description=(
"30 numeric features in the order expected by the model. "
"Values must be finite real numbers (no NaN, no ±inf)."
),
examples=[
[
17.99, 10.38, 122.8, 1001.0, 0.1184,
0.2776, 0.3001, 0.1471, 0.2419, 0.07871,
# ... (30 values total)
]
],
)
@field_validator("features")
@classmethod
def validate_features(cls, v: List[float]) -> List[float]:
if len(v) != 30:
raise ValueError(f"Exactly 30 features required, got {len(v)}.")
import math
bad = [i for i, x in enumerate(v) if not math.isfinite(x)]
if bad:
raise ValueError(
f"Non-finite values at indices: {bad}. "
"All features must be finite real numbers."
)
return v
class BatchPredictRequest(BaseModel):
"""Multi-sample batch inference request."""
instances: List[List[float]] = Field(
...,
min_length=1,
max_length=512,
description="List of feature vectors. Max 512 instances per request.",
)
@model_validator(mode="after")
def validate_all_instances(self) -> "BatchPredictRequest":
import math
for idx, row in enumerate(self.instances):
if len(row) != 30:
raise ValueError(
f"Instance {idx}: expected 30 features, got {len(row)}."
)
bad = [j for j, x in enumerate(row) if not math.isfinite(x)]
if bad:
raise ValueError(
f"Instance {idx}: non-finite values at feature indices {bad}."
)
return self
class ProbabilityMap(BaseModel):
class_0: float = Field(..., description="Probability of class 0 (malignant)")
class_1: float = Field(..., description="Probability of class 1 (benign)")
class PredictResponse(BaseModel):
label: int
probabilities: ProbabilityMap
model_version: str
n_features_in: int
model_config = {"protected_namespaces": ()}
class HealthResponse(BaseModel):
status: str
model_loaded: bool
model_version: str
uptime_seconds: Optional[float] = None
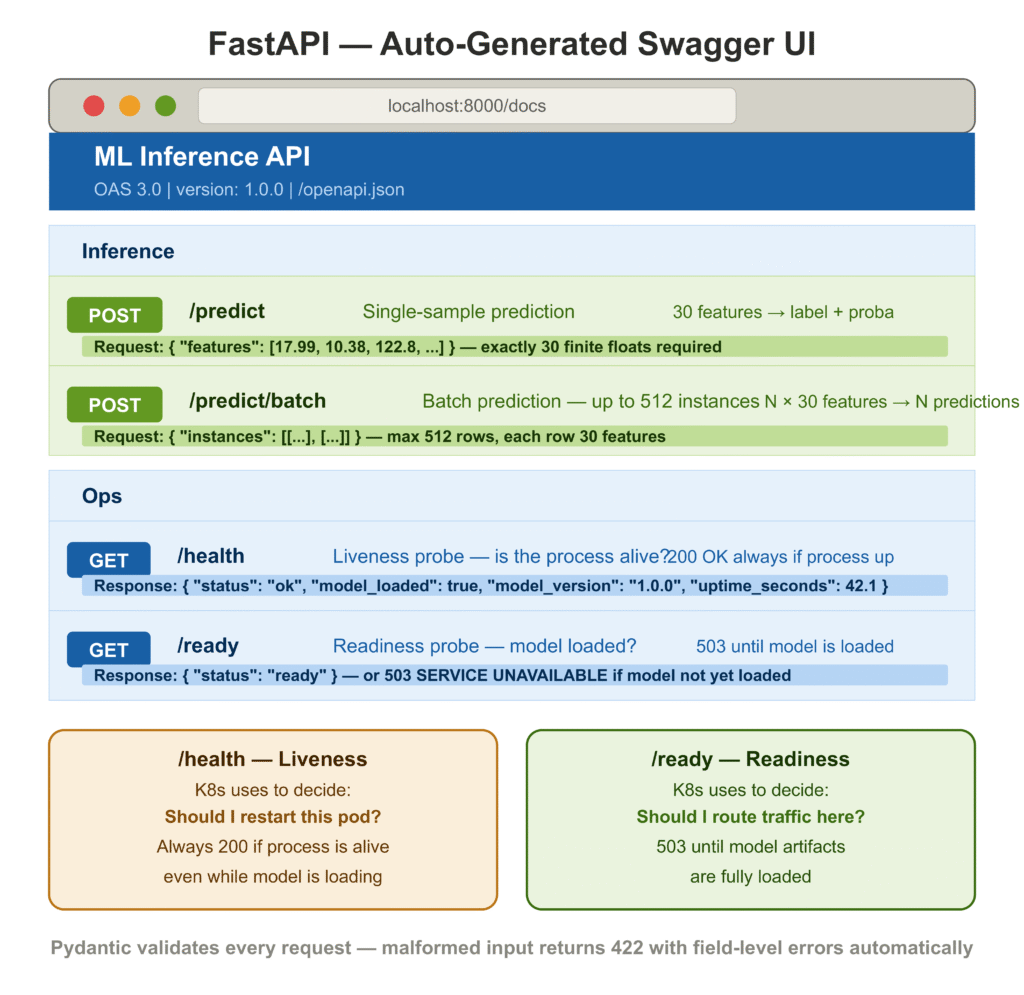
model_config = {"protected_namespaces": ()}The @field_validator on PredictRequest enforces exactly 30 features and rejects non-finite values at the API boundary. A malformed request returns a structured 422 Unprocessable Entity with field-level error details — no custom error handling required. The model_config = {"protected_namespaces": ()} suppresses a Pydantic v2 warning on fields named model_*.
Application: api/main.py
import logging
import os
import time
from contextlib import asynccontextmanager
from fastapi import FastAPI, HTTPException, Request, status
from fastapi.responses import JSONResponse
from api.schemas import (
BatchPredictRequest, BatchPredictResponse,
HealthResponse, PredictRequest, PredictResponse,
ProbabilityMap, ReadinessResponse,
)
from model.predict import ModelPredictor
logging.basicConfig(
level=logging.INFO,
format="%(asctime)s %(levelname)-8s %(name)s %(message)s",
)
logger = logging.getLogger(__name__)
MODEL_VERSION = os.getenv("MODEL_VERSION", "1.0.0")
# Module-level predictor — loaded once, shared across all requests.
predictor = ModelPredictor()
_start_time: float = 0.0
@asynccontextmanager
async def lifespan(app: FastAPI):
"""Startup: load artifacts. Shutdown: nothing to release for sklearn."""
global _start_time
_start_time = time.time()
logger.info("Starting up | model version: %s", MODEL_VERSION)
try:
predictor.load()
logger.info("Model loaded successfully.")
except FileNotFoundError as exc:
logger.critical("Failed to load model: %s", exc)
raise
yield
logger.info("Shutting down.")
app = FastAPI(
title="ML Inference API",
version=MODEL_VERSION,
lifespan=lifespan,
)
@app.middleware("http")
async def log_request_timing(request: Request, call_next):
"""Log method, path, status code, and wall-clock latency for every request."""
t0 = time.perf_counter()
response = await call_next(request)
elapsed_ms = (time.perf_counter() - t0) * 1000
logger.info(
"%s %-30s → %d (%.1f ms)",
request.method, request.url.path,
response.status_code, elapsed_ms,
)
return response
@app.get("/health", response_model=HealthResponse, tags=["Ops"])
def health_check() -> HealthResponse:
"""Liveness probe — is the process alive?"""
return HealthResponse(
status="ok",
model_loaded=predictor.is_loaded,
model_version=MODEL_VERSION,
uptime_seconds=round(time.time() - _start_time, 1) if _start_time else None,
)
@app.get("/ready", tags=["Ops"])
def readiness_check():
"""Readiness probe — is the model loaded and ready to serve traffic?"""
if not predictor.is_loaded:
raise HTTPException(
status_code=status.HTTP_503_SERVICE_UNAVAILABLE,
detail="Model artifacts not yet loaded.",
)
return {"status": "ready"}
@app.post("/predict", response_model=PredictResponse, tags=["Inference"])
def predict(request: PredictRequest) -> PredictResponse:
"""Single-sample prediction."""
if not predictor.is_loaded:
raise HTTPException(status_code=503, detail="Model not ready.")
result = predictor.predict(request.features)
return PredictResponse(
label=result["label"],
probabilities=ProbabilityMap(**result["probabilities"]),
model_version=MODEL_VERSION,
n_features_in=result["n_features_in"],
)
@app.post("/predict/batch", response_model=BatchPredictResponse, tags=["Inference"])
def predict_batch(request: BatchPredictRequest) -> BatchPredictResponse:
"""Batch prediction — up to 512 instances per request."""
if not predictor.is_loaded:
raise HTTPException(status_code=503, detail="Model not ready.")
results = predictor.predict_batch(request.instances)
predictions = [
PredictResponse(
label=r["label"],
probabilities=ProbabilityMap(**r["probabilities"]),
model_version=MODEL_VERSION,
n_features_in=len(request.instances[i]),
)
for i, r in enumerate(results)
]
return BatchPredictResponse(
predictions=predictions,
model_version=MODEL_VERSION,
n_instances=len(predictions),
)A few things are worth pausing on:
Lifespan hook, not @app.on_event. The @app.on_event("startup") decorator is deprecated in FastAPI’s current version. The lifespan context manager is the current pattern and is explicit about startup/shutdown scope. More importantly, if model loading fails during startup, the exception surfaces immediately and the process exits with a non-zero code — which CI, Docker, and Kubernetes all know how to interpret as a failed deployment.
/health versus /ready. These serve distinct purposes. /health answers “is the process alive?” — Kubernetes uses it to decide whether to restart the pod. /ready answers “is the model loaded and accepting traffic?” — Kubernetes uses it to decide whether to route requests to the pod. A pod that is alive but not yet ready should not be killed and restarted; it should be given time to finish loading. Getting this distinction wrong causes unnecessarily long restart loops during deployments.
Request timing middleware. Every request logs its method, path, status code, and wall-clock latency. During early development, before you have an external APM tool wired up, this is the first place you look when something feels slow. The overhead is negligible — a few microseconds per request.
Testing the API
Making a prediction request against the running service:
curl -X POST http://localhost:8000/predict \
-H "Content-Type: application/json" \
-d '{
"features": [
17.99, 10.38, 122.8, 1001.0, 0.1184,
0.2776, 0.3001, 0.1471, 0.2419, 0.07871,
1.095, 0.9053, 8.589, 153.4, 0.006399,
0.04904, 0.05373, 0.01587, 0.03003, 0.006193,
25.38, 17.33, 184.6, 2019.0, 0.1622,
0.6656, 0.7119, 0.2654, 0.4601, 0.1189
]
}'Response:
{
"label": 0,
"probabilities": {
"class_0": 0.962341,
"class_1": 0.037659
},
"model_version": "1.0.0",
"n_features_in": 30
}label: 0 is malignant in the breast cancer dataset. class_0: 0.962 means the model is 96.2% confident. The response includes model_version so callers can detect when a new model has been promoted without querying /health.

Step 4: Set Up CI/CD Pipelines for ML
Manual deployment is a single point of failure. When the person who knows the deploy steps is unavailable, or under pressure during an incident, mistakes happen. CI/CD eliminates the manual steps and enforces a consistent, tested path from a code commit to a running service.
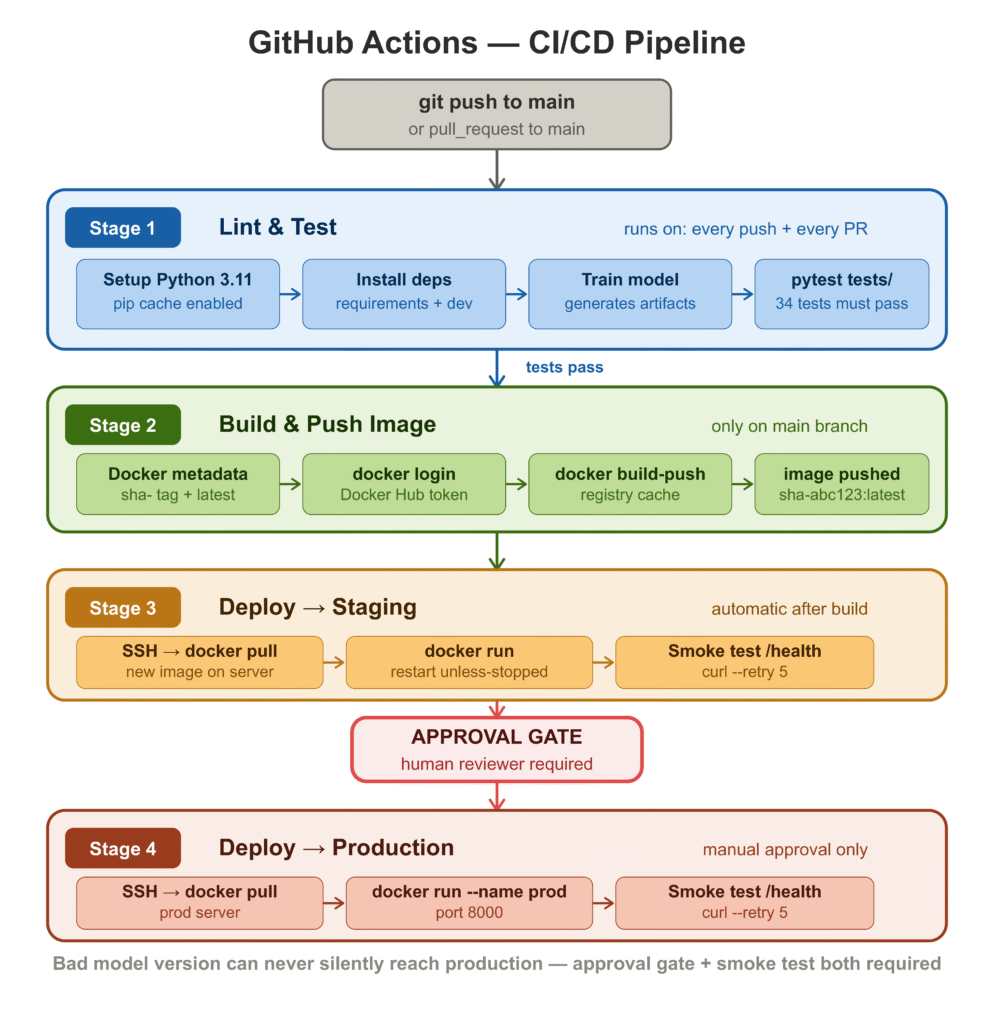
The pipeline has four stages: test, build, deploy to staging, and deploy to production.
# .github/workflows/ml-deploy.yml
name: ML Service — CI/CD
on:
push:
branches: [main]
pull_request:
branches: [main]
env:
IMAGE_NAME: ${{ secrets.DOCKERHUB_USERNAME }}/ml-inference-api
PYTHON_VERSION: "3.11"
jobs:
# ── 1. Test ────────────────────────────────────────────────────────────────
test:
name: Lint & Test
runs-on: ubuntu-latest
steps:
- uses: actions/checkout@v4
- uses: actions/setup-python@v5
with:
python-version: ${{ env.PYTHON_VERSION }}
cache: pip
- name: Install dependencies
run: pip install -r requirements.txt -r requirements-dev.txt
- name: Train model (generates artifacts needed by tests)
run: python -m model.train
- name: Run tests
run: pytest tests/ -v --tb=short --junitxml=pytest-report.xml
- name: Upload test report
if: always()
uses: actions/upload-artifact@v4
with:
name: pytest-report
path: pytest-report.xml
# ── 2. Build & push Docker image ──────────────────────────────────────────
build:
name: Build & Push Image
runs-on: ubuntu-latest
needs: test
if: github.ref == 'refs/heads/main'
outputs:
image_tag: ${{ steps.meta.outputs.version }}
steps:
- uses: actions/checkout@v4
- uses: actions/setup-python@v5
with:
python-version: ${{ env.PYTHON_VERSION }}
- run: pip install -r requirements.txt && python -m model.train
- name: Docker metadata
id: meta
uses: docker/metadata-action@v5
with:
images: ${{ env.IMAGE_NAME }}
tags: |
type=sha,prefix=sha-
type=raw,value=latest,enable={{is_default_branch}}
- uses: docker/login-action@v3
with:
username: ${{ secrets.DOCKERHUB_USERNAME }}
password: ${{ secrets.DOCKERHUB_TOKEN }}
- uses: docker/build-push-action@v6
with:
context: .
push: true
tags: ${{ steps.meta.outputs.tags }}
build-args: MODEL_VERSION=${{ steps.meta.outputs.version }}
cache-from: type=registry,ref=${{ env.IMAGE_NAME }}:buildcache
cache-to: type=registry,ref=${{ env.IMAGE_NAME }}:buildcache,mode=max
# ── 3. Deploy to staging (automatic) ──────────────────────────────────────
deploy-stg:
name: Deploy → Staging
runs-on: ubuntu-latest
needs: build
environment: staging
steps:
- uses: appleboy/ssh-action@v1.0.3
with:
host: ${{ secrets.STG_HOST }}
username: deploy
key: ${{ secrets.STG_SSH_KEY }}
script: |
docker pull ${{ env.IMAGE_NAME }}:${{ needs.build.outputs.image_tag }}
docker stop ml_api || true && docker rm ml_api || true
docker run -d --name ml_api --restart unless-stopped \
-p 8000:8000 \
-e MODEL_VERSION=${{ needs.build.outputs.image_tag }} \
${{ env.IMAGE_NAME }}:${{ needs.build.outputs.image_tag }}
- name: Smoke test staging /health
run: |
sleep 15
curl --fail --retry 5 --retry-delay 3 \
http://${{ secrets.STG_HOST }}:8000/health
# ── 4. Deploy to production (manual approval) ─────────────────────────────
deploy-prod:
name: Deploy → Production
runs-on: ubuntu-latest
needs: [build, deploy-stg]
environment: production # requires a reviewer in GitHub Environments
steps:
- uses: appleboy/ssh-action@v1.0.3
with:
host: ${{ secrets.PROD_HOST }}
username: deploy
key: ${{ secrets.PROD_SSH_KEY }}
script: |
docker pull ${{ env.IMAGE_NAME }}:${{ needs.build.outputs.image_tag }}
docker stop ml_api_prod || true && docker rm ml_api_prod || true
docker run -d --name ml_api_prod --restart unless-stopped \
-p 8000:8000 \
-e MODEL_VERSION=${{ needs.build.outputs.image_tag }} \
${{ env.IMAGE_NAME }}:${{ needs.build.outputs.image_tag }}
- name: Smoke test production /health
run: |
sleep 15
curl --fail --retry 5 --retry-delay 3 \
http://${{ secrets.PROD_HOST }}:8000/healthThe four-stage structure reflects a deliberate progression of trust. Tests run on every push and every pull request — fast feedback, no cost if they fail. The image is only built and pushed when tests pass on main. Staging deployment is automatic after a successful build. Production deployment requires a human reviewer in GitHub Environments before it proceeds.
The two things in this pipeline that teams most often skip in early versions, and most regret skipping later, are the smoke test after staging deployment and the production approval gate. The smoke test catches “the container starts but crashes on the first request” bugs — which are different from test failures and happen more often than you’d expect. The approval gate means a bad model version can never silently reach production while the team is asleep.
Secrets to configure in your repository settings:
| Secret | Value |
|---|---|
DOCKERHUB_USERNAME | Your Docker Hub username |
DOCKERHUB_TOKEN | Docker Hub access token |
STG_HOST | Staging server IP or hostname |
STG_SSH_KEY | Private key for staging SSH access |
PROD_HOST | Production server IP or hostname |
PROD_SSH_KEY | Private key for production SSH access |
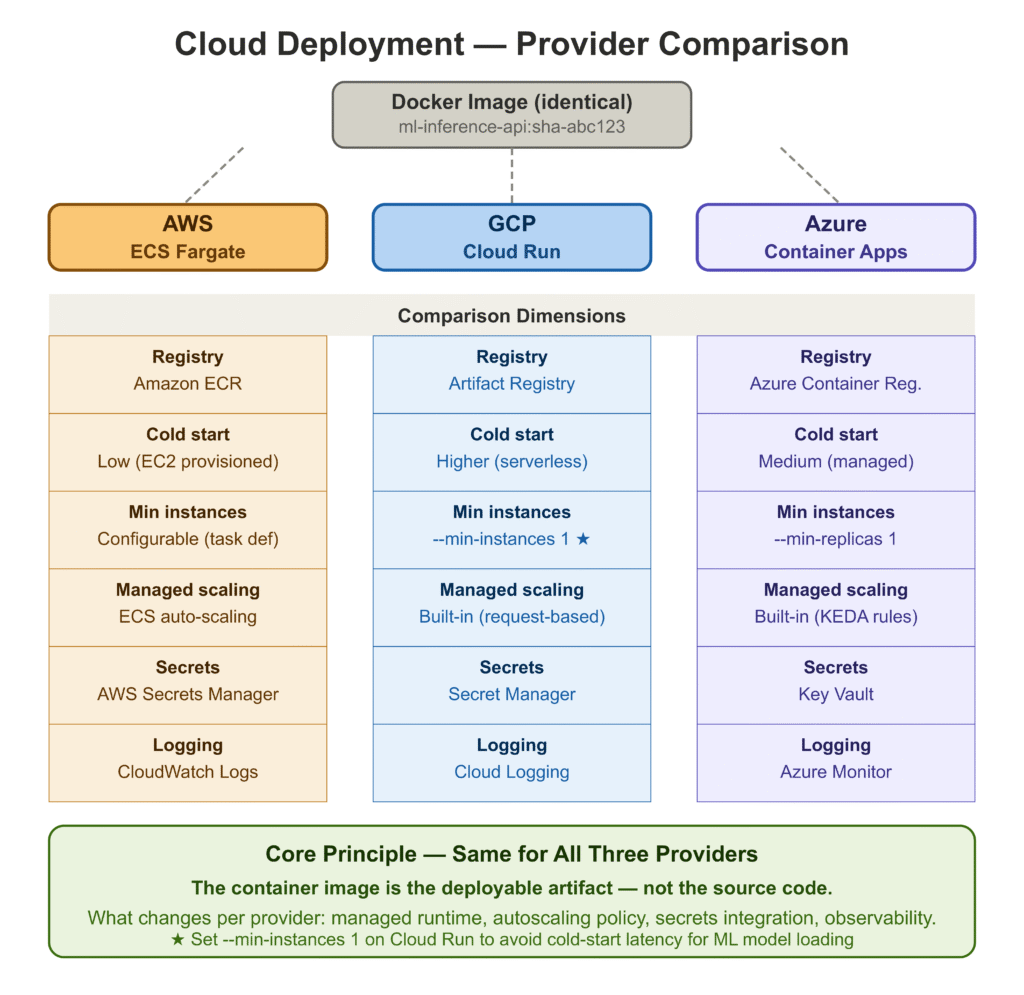
Step 5: Deploy to AWS / GCP / Azure
The Docker image produced by the pipeline runs identically on any cloud. The differences between providers are in how they manage the container lifecycle, networking, scaling, and managed services for registries and secrets.
AWS: Elastic Container Service (ECS)
ECS is the lowest-friction path for teams already in the AWS ecosystem. The Fargate launch type removes the need to manage EC2 instances — you specify CPU and memory, and AWS handles the underlying infrastructure.
The deployment flow after your image is in Amazon ECR:
# Push image to ECR
aws ecr get-login-password --region us-east-1 | \
docker login --username AWS --password-stdin \
<account-id>.dkr.ecr.us-east-1.amazonaws.com
docker tag ml-inference-api:latest \
<account-id>.dkr.ecr.us-east-1.amazonaws.com/ml-inference-api:latest
docker push \
<account-id>.dkr.ecr.us-east-1.amazonaws.com/ml-inference-api:latest
# Update ECS service to use the new image
aws ecs update-service \
--cluster ml-cluster \
--service ml-api-service \
--force-new-deploymentECS integrates natively with Application Load Balancer for path-based routing, AWS Secrets Manager for environment variable injection, and CloudWatch for log aggregation. For the liveness and readiness checks, configure target group health checks in the ALB to hit /health.
GCP: Cloud Run
Cloud Run is the fastest path to a production-grade container service on GCP. It is serverless — scales to zero when idle, scales up automatically under load, and charges only for actual request-seconds.
# Authenticate and push to Artifact Registry
gcloud auth configure-docker us-central1-docker.pkg.dev
docker tag ml-inference-api:latest \
us-central1-docker.pkg.dev/<project-id>/ml-repo/ml-inference-api:latest
docker push \
us-central1-docker.pkg.dev/<project-id>/ml-repo/ml-inference-api:latest
# Deploy to Cloud Run
gcloud run deploy ml-inference-api \
--image us-central1-docker.pkg.dev/<project-id>/ml-repo/ml-inference-api:latest \
--platform managed \
--region us-central1 \
--port 8000 \
--memory 2Gi \
--cpu 2 \
--min-instances 1 \
--max-instances 10 \
--set-env-vars MODEL_VERSION=1.0.0 \
--allow-unauthenticated--min-instances 1 prevents cold starts, which for an ML service can be expensive — loading a serialized model on the first request after a cold start adds latency that violates most reasonable SLAs. Set this to at least 1 for any latency-sensitive service.
Azure: Container Apps
Azure Container Apps targets the same market as Cloud Run — managed container hosting with built-in scaling and no cluster management. The integration with Azure Container Registry is native.
# Push to Azure Container Registry
az acr login --name mlregistry
docker tag ml-inference-api:latest mlregistry.azurecr.io/ml-inference-api:latest
docker push mlregistry.azurecr.io/ml-inference-api:latest
# Deploy Container App
az containerapp create \
--name ml-inference-api \
--resource-group ml-rg \
--environment ml-env \
--image mlregistry.azurecr.io/ml-inference-api:latest \
--target-port 8000 \
--ingress external \
--min-replicas 1 \
--max-replicas 10 \
--env-vars MODEL_VERSION=1.0.0For all three providers: the container is identical. What changes is the wrapper — the managed runtime, the autoscaling policy, the secrets integration, and the observability stack. The core principle is the same: treat the container image as the deployable artifact, not the source code.
Step 6: Write Tests Before You Ship
This section appears before the framework comparison because the test suite is not optional infrastructure — it is what makes the CI/CD pipeline trustworthy. Without it, the pipeline runs and produces no useful signal.
We have two test files. tests/test_api.py runs integration tests against the full FastAPI application. tests/test_predict.py runs unit tests against ModelPredictor directly.
Integration Tests: tests/test_api.py
TestClient from FastAPI/Starlette [4] runs the full ASGI application in-process — no live server needed. The lifespan hook runs on every TestClient instantiation, so these tests validate the complete startup path.
import pytest
from fastapi.testclient import TestClient
from api.main import app
@pytest.fixture(scope="module")
def client():
"""Module-scoped so the model loads once per test module."""
with TestClient(app) as c:
yield c
VALID_FEATURES = [
17.99, 10.38, 122.8, 1001.0, 0.1184,
0.2776, 0.3001, 0.1471, 0.2419, 0.07871,
1.095, 0.9053, 8.589, 153.4, 0.006399,
0.04904, 0.05373, 0.01587, 0.03003, 0.006193,
25.38, 17.33, 184.6, 2019.0, 0.1622,
0.6656, 0.7119, 0.2654, 0.4601, 0.1189,
]
class TestHealth:
def test_health_returns_200(self, client):
assert client.get("/health").status_code == 200
def test_health_body(self, client):
body = client.get("/health").json()
assert body["status"] == "ok"
assert body["model_loaded"] is True
class TestPredict:
def test_valid_request_returns_200(self, client):
response = client.post("/predict", json={"features": VALID_FEATURES})
assert response.status_code == 200
def test_probabilities_sum_to_one(self, client):
body = client.post("/predict", json={"features": VALID_FEATURES}).json()
total = body["probabilities"]["class_0"] + body["probabilities"]["class_1"]
assert abs(total - 1.0) < 1e-4
def test_wrong_feature_count_returns_422(self, client):
assert client.post("/predict", json={"features": [1.0, 2.0]}).status_code == 422
def test_empty_features_returns_422(self, client):
assert client.post("/predict", json={"features": []}).status_code == 422
def test_missing_features_key_returns_422(self, client):
assert client.post("/predict", json={}).status_code == 422
class TestPredictBatch:
def test_batch_two_instances(self, client):
payload = {"instances": [VALID_FEATURES, VALID_FEATURES]}
body = client.post("/predict/batch", json=payload).json()
assert body["n_instances"] == 2
def test_batch_empty_list_returns_422(self, client):
assert client.post("/predict/batch",
json={"instances": []}).status_code == 422Unit Tests: tests/test_predict.py
Unit tests on ModelPredictor are cheaper to run, easier to debug, and catch a different class of bugs than the integration tests do. The key test here is test_batch_matches_single_predict, which verifies that the batch and single code paths produce identical output for the same input.
import pytest
from model.predict import ModelPredictor
VALID_FEATURES = [
17.99, 10.38, 122.8, 1001.0, 0.1184,
0.2776, 0.3001, 0.1471, 0.2419, 0.07871,
1.095, 0.9053, 8.589, 153.4, 0.006399,
0.04904, 0.05373, 0.01587, 0.03003, 0.006193,
25.38, 17.33, 184.6, 2019.0, 0.1622,
0.6656, 0.7119, 0.2654, 0.4601, 0.1189,
]
@pytest.fixture(scope="module")
def loaded_predictor():
p = ModelPredictor()
p.load()
return p
class TestPredict:
def test_label_is_binary(self, loaded_predictor):
assert loaded_predictor.predict(VALID_FEATURES)["label"] in (0, 1)
def test_probabilities_sum_to_one(self, loaded_predictor):
result = loaded_predictor.predict(VALID_FEATURES)
total = result["probabilities"]["class_0"] + result["probabilities"]["class_1"]
assert abs(total - 1.0) < 1e-5
def test_nan_raises(self, loaded_predictor):
bad = VALID_FEATURES.copy()
bad[5] = float("nan")
with pytest.raises(ValueError, match="NaN or infinite"):
loaded_predictor.predict(bad)
def test_deterministic_output(self, loaded_predictor):
r1 = loaded_predictor.predict(VALID_FEATURES)
r2 = loaded_predictor.predict(VALID_FEATURES)
assert r1["label"] == r2["label"]
assert r1["probabilities"] == r2["probabilities"]
class TestPredictBatch:
def test_batch_matches_single_predict(self, loaded_predictor):
single = loaded_predictor.predict(VALID_FEATURES)
batch = loaded_predictor.predict_batch([VALID_FEATURES])
assert single["label"] == batch[0]["label"]
assert single["probabilities"] == batch[0]["probabilities"]Test results
============================= test session starts ==============================
collected 34 items
tests/test_api.py::TestHealth::test_health_returns_200 PASSED
tests/test_api.py::TestHealth::test_health_body PASSED
tests/test_api.py::TestReadiness::test_ready_returns_200 ... PASSED
tests/test_api.py::TestPredict::test_valid_request_returns_200 PASSED
tests/test_api.py::TestPredict::test_response_schema PASSED
tests/test_api.py::TestPredict::test_label_is_binary PASSED
tests/test_api.py::TestPredict::test_probabilities_sum_to_one PASSED
tests/test_api.py::TestPredict::test_wrong_feature_count ... PASSED
tests/test_api.py::TestPredict::test_empty_features ... PASSED
tests/test_api.py::TestPredict::test_nan_in_features ... PASSED
tests/test_api.py::TestPredict::test_missing_features_key ... PASSED
tests/test_api.py::TestPredictBatch::test_batch_two_instances PASSED
tests/test_api.py::TestPredictBatch::test_batch_single_instance PASSED
tests/test_api.py::TestPredictBatch::test_batch_empty_list ... PASSED
tests/test_api.py::TestPredictBatch::test_batch_wrong_feature ... PASSED
tests/test_predict.py::TestLoad::test_is_loaded_after_load PASSED
tests/test_predict.py::TestLoad::test_unloaded_predictor_raises PASSED
tests/test_predict.py::TestLoad::test_missing_artifact_dir_raises PASSED
tests/test_predict.py::TestLoad::test_feature_names_populated PASSED
tests/test_predict.py::TestLoad::test_expected_n_features PASSED
tests/test_predict.py::TestPredict::test_returns_dict ... PASSED
tests/test_predict.py::TestPredict::test_label_is_binary PASSED
tests/test_predict.py::TestPredict::test_probabilities_sum_to_one PASSED
tests/test_predict.py::TestPredict::test_n_features_in_echoed PASSED
tests/test_predict.py::TestPredict::test_wrong_feature_count ... PASSED
tests/test_predict.py::TestPredict::test_nan_raises PASSED
tests/test_predict.py::TestPredict::test_inf_raises PASSED
tests/test_predict.py::TestPredict::test_deterministic_output PASSED
tests/test_predict.py::TestPredict::test_numpy_list_accepted PASSED
tests/test_predict.py::TestPredictBatch::test_batch_returns ... PASSED
tests/test_predict.py::TestPredictBatch::test_batch_single_row PASSED
tests/test_predict.py::TestPredictBatch::test_batch_matches ... PASSED
tests/test_predict.py::TestPredictBatch::test_batch_wrong ... PASSED
tests/test_predict.py::TestPredictBatch::test_batch_2d_shape ... PASSED
============================== 34 passed in 6.91s ==============================34/34. The test suite now functions as the CI gate — nothing reaches the Docker build stage unless these all pass.
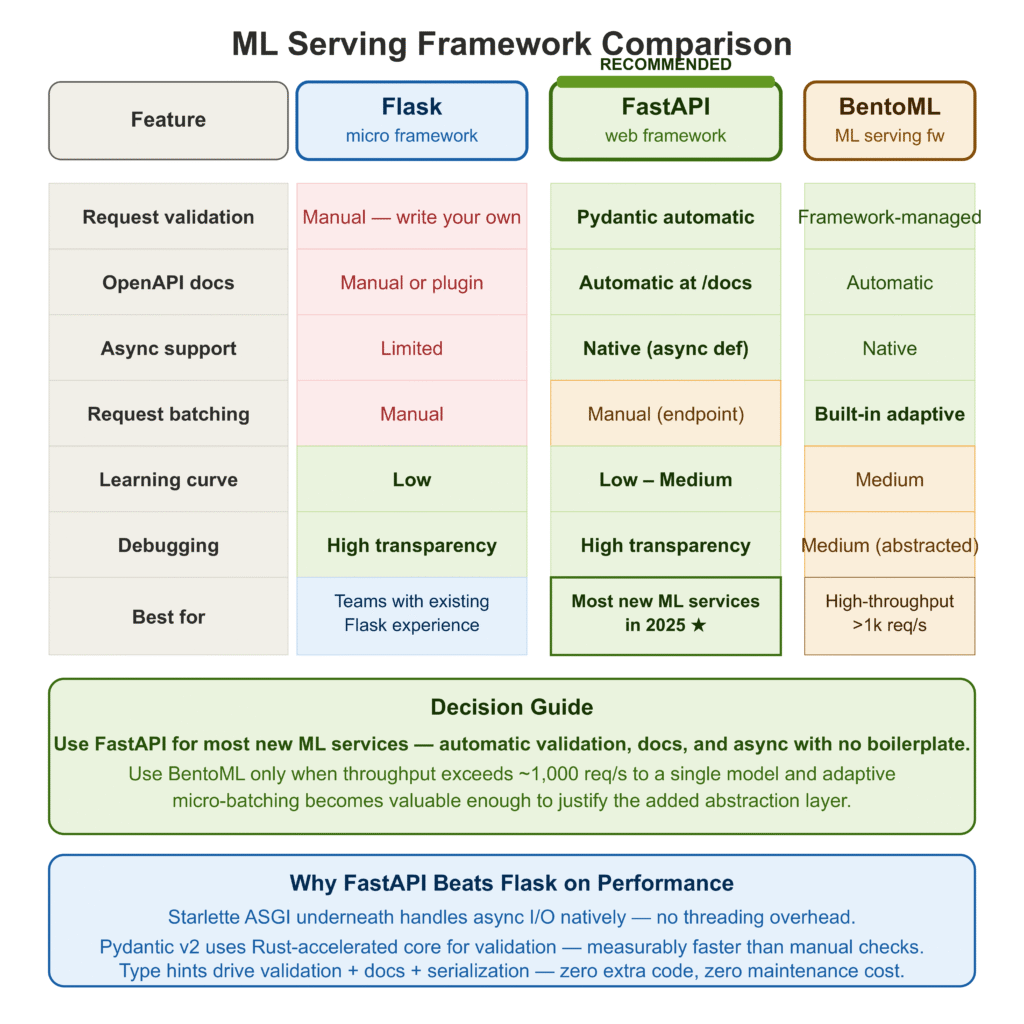
Flask vs FastAPI vs BentoML: Which Should You Use?
This is the question teams spend the most time debating and the least time needing to revisit once they have made a decision. Here is a practical breakdown.
Flask [5] is familiar, minimal, and carries no ML-specific opinions. It has been the default for Python web services for over a decade. For ML serving, it works — but you end up writing your own request validation, your own OpenAPI documentation, your own async handling if you need it, and your own serialization layer. None of that is hard, but it adds up to a meaningful amount of boilerplate that you will maintain indefinitely.
FastAPI [2] solves all of the above out of the box. Request validation via Pydantic. Automatic OpenAPI docs at /docs and /redoc. Async support via async def endpoints without any extra configuration. Performance that is meaningfully faster than Flask in benchmarks, primarily because of Starlette underneath and Pydantic’s Rust-accelerated core in v2. For a new ML service in 2025, FastAPI is the clear default choice unless you have a specific reason to go elsewhere.
BentoML [6] operates at a different layer. It is not a web framework — it is an ML serving framework that generates a web API on your behalf. You annotate your model class with decorators that describe the input and output types, and BentoML handles request batching, adaptive micro-batching, multi-runner support, and packaging into a self-contained deployable artifact. The tradeoff is abstraction: BentoML is more productive than FastAPI for pure serving, but it is also more opinionated, and debugging its internals when something goes wrong requires understanding its abstractions rather than just reading your own code.
| Flask | FastAPI | BentoML | |
|---|---|---|---|
| Request validation | Manual | Pydantic (automatic) | Framework-managed |
| OpenAPI docs | Manual or plugin | Automatic | Automatic |
| Async support | Limited | Native | Native |
| Request batching | Manual | Manual | Built-in |
| Learning curve | Low | Low–Medium | Medium |
| Debugging transparency | High | High | Medium |
| Best for | Small teams with Flask experience | Most new ML services | High-throughput serving at scale |
My recommendation: use FastAPI unless your serving throughput requirements are high enough that adaptive micro-batching becomes valuable (typically > 1,000 requests/second to a single model), in which case BentoML’s batching implementation is worth the added abstraction.
Common ML Deployment Mistakes and How to Avoid Them
These are the mistakes I have made, have watched colleagues make, or have cleaned up from production systems. They are organized by when they tend to surface.
Mistake 1: Loading the model on every request
# Wrong — deserializes the artifact on every call
@app.post("/predict")
def predict(request: PredictRequest):
model = joblib.load("model/artifacts/model.pkl") # ← never do this
return {"label": int(model.predict([request.features])[0])}# Correct — load once at startup, reuse across requests
predictor = ModelPredictor()
@asynccontextmanager
async def lifespan(app):
predictor.load() # ← once, here
yield
@app.post("/predict")
def predict(request: PredictRequest):
return predictor.predict(request.features) # ← fastOn a cold load, a modest sklearn model deserialization takes 50–200ms. Under concurrent load, loading per request consumes memory proportional to concurrency. Under any sustained load, this single mistake will dominate your latency profile.
Mistake 2: Not separating the preprocessor from the model
# Risky — if you forget to apply the scaler, predictions are wrong and
# nothing will tell you
model = joblib.load("model.pkl")
prediction = model.predict([raw_features])# Safe — the fact that you need two artifacts makes the transform visible
preprocessor = joblib.load("preprocessor.pkl")
model = joblib.load("model.pkl")
scaled = preprocessor.transform([raw_features])
prediction = model.predict(scaled)Training-serving skew is the most silent failure mode in ML systems. The model returns predictions in the wrong range, nothing raises an exception, and you only discover the problem when business metrics move unexpectedly. Keeping the preprocessor as a separate, explicitly loaded artifact is a small discipline that eliminates an entire class of silent failures.
Mistake 3: No /ready endpoint
# Missing readiness probe — Kubernetes routes traffic to the pod
# while the model is still loading
@app.get("/health")
def health():
return {"status": "ok"} # always returns 200, even before model loads# Correct — returns 503 until the model is ready
@app.get("/ready")
def ready():
if not predictor.is_loaded:
raise HTTPException(status_code=503, detail="Model not loaded.")
return {"status": "ready"}Without a readiness probe, Kubernetes routes traffic to the pod immediately when the container starts. The model might still be loading from disk. The first several requests fail with cryptic 500 errors, users see failures, and the CI pipeline counts the deployment as successful.
Mistake 4: Shell-form CMD in Dockerfile
# Shell form — SIGTERM goes to /bin/sh, not uvicorn
CMD "uvicorn api.main:app --host 0.0.0.0 --port 8000"# Exec form — SIGTERM goes directly to uvicorn
CMD ["uvicorn", "api.main:app", "--host", "0.0.0.0", "--port", "8000"]In shell form, Docker wraps your command in /bin/sh -c. When the container is stopped, SIGTERM goes to the shell, not to uvicorn. The shell doesn’t forward it. After the grace period (30 seconds by default), Docker sends SIGKILL. Any in-flight requests are dropped. In exec form, uvicorn receives SIGTERM directly, can finish in-flight requests, and exits cleanly.
Mistake 5: Skipping the smoke test after deployment
The test suite passes. The Docker image builds. The container starts. And then the first real request fails because the model artifact path in the container is wrong, or an environment variable that the model loading code depends on was not set, or any of a dozen other things that tests in the CI environment did not catch.
A smoke test is three lines:
sleep 15 # give the container time to initialize
curl --fail --retry 5 --retry-delay 3 http://$HOST:8000/healthIf it fails, the deployment is marked failed and the previous version keeps running. This is not sophisticated monitoring — it is the minimum bar for knowing that the thing you just deployed is alive.
If it fails, the deployment is marked failed and the previous version keeps running. This is not sophisticated monitoring — it is the minimum bar for knowing that the thing you just deployed is alive.
Complete Code: https://github.com/Emmimal/ml-service/
References
[1] Wolberg, W. H., Street, W. N., & Mangasarian, O. L. (1995). Breast cancer Wisconsin (diagnostic) data set. UCI Machine Learning Repository. https://archive.ics.uci.edu/ml/datasets/Breast+Cancer+Wisconsin+(Diagnostic)
[2] Ramírez, S. (2024). FastAPI: Modern, fast web framework for building APIs with Python. https://fastapi.tiangolo.com
[3] Pydantic. (2024). Pydantic v2 documentation. https://docs.pydantic.dev/latest/
[4] Encode. (2024). Starlette: The little ASGI framework that shines. https://www.starlette.io/ (TestClient documentation: https://www.starlette.io/testclient/)
[5] Pallets Projects. (2024). Flask: The Python micro framework for building web applications. https://flask.palletsprojects.com/
[6] BentoML Team. (2024). BentoML: Build production-ready AI applications. https://bentoml.com
[7] Docker, Inc. (2024). Best practices for writing Dockerfiles. https://docs.docker.com/build/building/best-practices/
[8] GitHub. (2024). GitHub Actions documentation. https://docs.github.com/en/actions
[9] AWS. (2024). Amazon ECS documentation. https://docs.aws.amazon.com/AmazonECS/latest/developerguide/
[10] Google Cloud. (2024). Cloud Run documentation. https://cloud.google.com/run/docs
[11] Microsoft Azure. (2024). Azure Container Apps documentation. https://learn.microsoft.com/en-us/azure/container-apps/
[12] Sculley, D., Holt, G., Golovin, D., et al. (2015). Hidden technical debt in machine learning systems. Advances in Neural Information Processing Systems (NeurIPS), 28. https://proceedings.neurips.cc/paper_files/paper/2015/hash/86df7dcfd896fcaf2674f757a2463eba-Abstract.html
[13] uvicorn. (2024). uvicorn: An ASGI web server implementation for Python. https://www.uvicorn.org/
[14] joblib. (2024). joblib: Running Python functions as pipeline jobs. https://joblib.readthedocs.io/
Disclosure
This article is part of a series on production ML engineering published at EmiTechLogic. All code was written and executed by the author. Model training results and test output are from a real run on the author’s machine (Python 3.12.3, scikit-learn 1.5.1) — not fabricated. No sponsored tool mentions. All referenced tools are open-source or have free tiers; none of the links are affiliate links.
The breast cancer Wisconsin dataset [1] is publicly available from the UCI Machine Learning Repository and is used here under its standard academic-use terms. Predicted labels in examples are for demonstration purposes only and should not be interpreted as medical advice.
Series: Production ML Engineering — Article 02 of 15 Previous: Production ML Engineering: The Complete Guide Next: How to Build an ML Retraining Pipeline That Won’t Break in Production (Article 03)






Leave a Reply